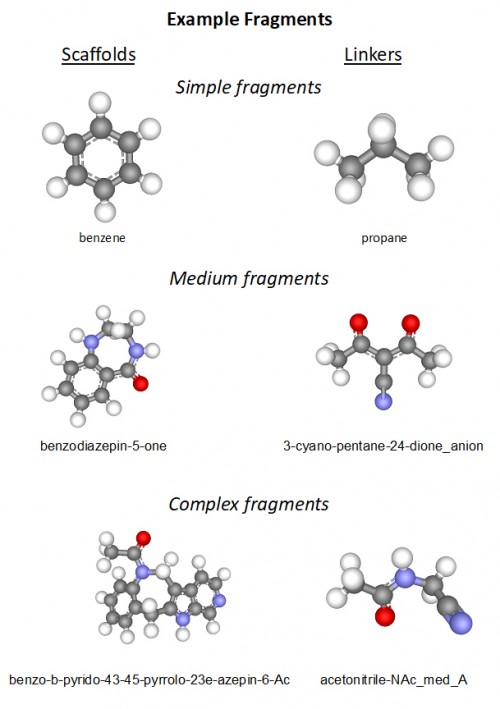
What Are Fragment Maps?
Fragment maps describe the binding of chemical fragments and waters to macromolecules such as proteins, RNA, and DNA. A chemical fragment is a small chemical component used to compose drug molecules. Our optimized Monte Carlo simulator (BMOC) is used to produce statistical distributions of fragment binding configurations on the surface of macromolecules. These distributions conform to rigorous Boltzmann energy statistics. The resulting fragment binding data consists of 100-500 samples (“snapshots”) of the 3-D location and 3-axis orientation of rigid fragments. Typically, each snapshot includes hundreds of individual fragments at different locations. A collection of snapshots is referred to as a fragment map. As many as several hundred different simulations are run on a protein, using diverse fragment types representing a range of chemical components, such as rings (e.g. pyrimidine), linkers (e.g. propane), and combinations (e.g. biphenyl). Users can also run their own fragment sets.
How many fragment maps have been computed?
We have currently simulated almost 500 proteins, with a goal of reaching several thousand. Currently, 136 fragment and water simulations are carried out for each protein at over 20 chemical potential annealing steps (the exact number varies depending on the fragment type). Typically, 100 samples or snapshots are taken for each annealing step, resulting in more than 300,000 samples per protein. Each sample has from 1-50,000 fragment poses. Thus, there are hundreds of millions of samples consisting, in aggregate, of billions of fragment poses. The fragments to simulate are selected from a library of ~3,500 fragments, and users can define their own fragments to be simulated incrementally on the proteins.
Why is this fragment binding data important?
Fragment binding data from BMOC simulations provides statistically significant orientations of fragment binding poses, in contrast to probing methods that produce individual poses which may or may not be statistically representative. This is important when search algorithms evaluate whether a given pose should be suggested – is it near the center of the distribution or on the fringe? Secondly, our fragment binding data is ranked by excess chemical potential, which, for isolated sites, is the free energy of binding [[Footnote: configuration free energy of binding]]. This ranking is critical for prioritizing potential fragment substitutions to a compound. By incorporating entropy, this ranking has been demonstrated to be reliable when protein flexibility is a small effect. The top-ranked fragment substitutions are “what the protein wants,” giving compounds a head start to better affinity
How are fragment maps made available?
Fragment maps represent an enormous amount of detailed information, very tedious to use directly. The BMaps web app incorporates a geometric search algorithm to interrogate the fragment binding data to find binding poses compatible with bond formation to an existing compound binding pose. The user just selects from a prioritized list of substitutions. This is different from other software which just presents fragments that can sterically fit.